Unincorporated dNTPs were removed in the probe with Illustra ProbeQuant G 50 Micro Columns from GE Healthcare. Hybridisation of BAC filters with MHC probes was carried out overnight at 60 C in Amersham Rapid hyb Buffer. Excess background and non distinct binding was mini mized by washing the filters at 60 C having a series of buffers. Amersham Hyperfilm MP was utilized to visualize positive clones after a single to 3 days of publicity for the filters. BAC clone sequencing BAC DNA of your optimistic clones was purified from a hundred ml LBchloramphenicol bacterial culture utilizing QIAGEN Huge Construct Kit. BACs containing distinctive MHC Class I or II genes were confirmed by direct end sequencing of BAC DNA with MHC primers, making use of normal sequencing support on the Australian Genome Exploration Facility Ltd.
Fingerprinting examination and comprehensive sequencing of MHC favourable BAC clones was conducted in the Wellcome Believe in Sanger Institute, Cambridge, United kingdom. Common Sanger sequencing kinase inhibitor mapk inhibitors approach was employed to ensure large assem bly accuracy of paralogous MHC genes. BAC sequence annotation BAC sequences have been aligned with human genomic and transcript sequences, non human reference RNA sequences, and identified or predicted opossum and tammar wallaby transcripts employing web based BLAST packages. Genes had been annotated manually determined by the top BLAST hits and in accordance using the suggestions from the Human and vertebrate analysis and annotation recommendations. Overlapping BAC sequences have been identified and aligned using BLASTN and ClustalW applications. A wallaby LINE 1 section was employed to hunt for putative LINE segments.
Fluorescent in situ hybridisation BAC clones containing MHC Class I or II genes had been physically mapped to a male devil karyotype selleckchem following the protocol described previously by Alsop and collea gues. Approximately one mg of BAC DNA was utilised to produce probes that have been labelled by nick translation with both SpectrumOrange dUTP or SpectrumGreen dUTP. Labelled probes had been hybridised overnight to devil chromosomes, which have been denatured for one min forty sec. Slides had been washed as soon as at 60 C in 0. 4x SSC with 0. 3% Tween20 for 2 min after which once in 2x SSC with 0. 1% Tween20 for 30 sec at space temperature. Chromosomes were counterstained in DAPI and mounted in VECTASHIELD Mounting Med ium from Vector Laboratories Inc. A Zeiss Axioplan2 epifluorescence microscope was utilised to visualize the fluorescent signals. Images of DAPI stained metaphase chromosomes and fluorescent signals were captured working with a SPOT RT Monochrome CCD charge coupled gadget camera and merged employing IP Lab imaging computer software. Sequencing and examination of MHC class I alleles To check the Class I gene number variation hypothesis, it had been necessary to ensure that all Class I loci in both Cedric and Spirit were characterized.
Monthly Archives: July 2014
005, no differentially expressed genes have been detected As bei
005, no differentially expressed genes have been detected. As a begin to analyse the expression data, Venn diagrams were manufactured to identify genes that were differentially expressed in HacACA at all three time factors when com pared on the wild form strain. As shown in Figure 3A, 616 genes had been up regulated while in the constitutive HacA strain in any way three time points and 433 genes have been down regulated. A full listing of all expression information and also the FDR values for that pair smart comparison in the distinctive strains and time factors is given in. In the 616 up regulated genes we have been able to retrieve 598 upstream areas. These upstream areas were analysed to the presence of UPRE sequences NTGTGCCT three. From the up regulated genes from the HacACA strain, we found 47 genes that contained no less than one particular UPRE sequence inside the 400 bp area up stream their start codon.
In contrast towards the frequency of UPRE within the 400 bp up stream area of the remaining non up regulated genes a statistical important enrichment was assessed using the Fishers precise check. Whilst this analysis signifies a statistical enrichment for genes containing a HacA binding web site while in the promoter region of HacA induced genes, it demonstrates that only about 10% of selleck chemicals NU7441 the HacACA induced genes have a putative HacA binding internet site. It suggests that either the at the moment utilised HacA binding consensus web-site is as well stringent and that add itional sequences permit HacA to bind, or that further transcription aspects are involved during the induction in response to the constitutive activation of HacA.
The data set of HacA induced genes that has a putative UPRE web page involve genes associated to protein folding, lipid metabolism, transport inside the cell, glycosylation, ER high quality handle as well being a massive set of genes that code for hypothetical and unknown function proteins. Identification of biological processes enriched within the transcriptomic PD0332991 profiles on the HacACA strain To obtain an overview from the processes impacted in the transcriptional level among the HacAWT as well as HacACA one mutant, overrepresented GO terms amongst differentially expressed genes had been recognized. For this evaluation, we applied the Fishers precise check Gene Ontology annotation tool. Network maps of relevant GO terms , over or below represented within the HacACA strain, are offered in Add itional file six and Further file seven. In More file eight and Supplemental file 9, the results on the GO enrichment evaluation are provided.
To analyse the outcomes, two comple mentary approaches were taken. Firstly, we rationally defined GO terms of increased buy that contain many GO terms. Secondly, we looked exclusively at GO terms that are terminal during the network, as these annotations would be the most comprehensive. These approaches enabled us to determine four main categories of genes to describe probably the most related up regulated bio logical processes from the HacACA strain.
Promoter analysis for putative transcription factor binding sites
Promoter analysis for putative transcription factor binding sites Upstream regions proximal to the transcriptional start site of the rat Col2a1 and Agc1 genes have been described previ ously. Upstream regions from the transcriptional start site of the Rattus Norvegicus Col2a1 and Agc1 genes were obtained and analysed for putative transcription factor binding sites by TRANSFAC anal ysis. Oligodeoxynucleotide decoy assay Chondrocytes were plated at 1. 2106 cellswell in six well culture dishes. Single stranded, phosphorothiol modified ODNs were annealed by heating complementary ODNs to 98 C for 20 minutes followed by cooling to room temperature for 3 to 4 hours. Chondrocytes were transfected with 2M double stranded ODNs corresponding to the cognate EGR 1 binding sequence using 1% HiPerfect transfection reagent, as per the manufacturers instructions.
To optimize double stranded ODN transfection conditions, chondrocytes selleck chemicals were transfected cells with increasing concen trations of double stranded, fluorescein tagged and phospho rothiol modified ODNs, and the cells were imaged by live cell fluorescent microscopy. Chondrocytes were allowed to grow for 24 hours in the presence of ODNs, after which cells were washed and cultured in serum free RPMI media overnight. Chondrocytes were treated with TNF for 24 hours, as described, and total RNA was collected for analysis by real time PCR. Results ERK12 is phosphorylated by TNF in chondrocytes We have shown previously that TNF induces ERK phospho rylation in primary articular chondrocytes 15 minutes post treatment.
To confirm and extend these results, we used western blot analysis to show that TNF induced ERK12 phosphorylation 15 minutes post treatment, fol lowed by a dig this decrease in phosphorylation status. ERK12 phosphorylation was again increased at 90 minutes post treatment. As anticipated, both the increases at 15 minutes and at 90 minutes could be inhibited by the MEK12 inhibitor U0126, but not its inactive isoform U0124. Based on these data, we used U0126 as an inhibitor to assess the effect of blocking MEK12 on the mRNA expression pattern modulated by application of TNF to chondrocytes. U0126 blocks part of the TNF dependent gene expression changes in chondrocytes To investigate the global impact of U0126 on TNF modu lated gene expression in chondrocytes, we utilized microarrays to analyse changes in chondrocyte mRNA expression. Cells were serum starved overnight and were treated with or with out U0126 prior to addition of TNF for 24 hours. Cells were treated with TNF for 24 hours as previous data showed that this length of TNF treatment was neces sary to generate a TNF mediated suppression of chondro cyte matrix genes, owing to the stability of chondrocyte matrix gene mRNAs.
The primary report of IL 32 advocated that IL 32a stimulated TNFa
The primary report of IL 32 advocated that IL 32a stimulated TNFa manufacturing via activation of NF B and p38 MAPK in mouse RAW 267. four cells. the two peaks of p38 MAPK phosphorylation at five and 45 min utes had been regarded a characteristic choosing for this cytokine. Netea and colleagues subsequently reported that IL 32 induced TNFa production by human peripheral blood mononuclear cells was similarly regulated as a result of phosphorylation of p38 MAPK. Alternatively, ERK12 MAPK was domi nant in IL 32 induced osteoclastogenesis for human PBMCs and in IL 32 induced IL 6 and IL 8 pro duction by human fibroblast like synoviocytes. The current study demonstrated that IL 32a induced TNFa production was mediated through phosphorylation of I B and ERK12 in RAW 267. four cells.
Really, all 3 components of MAPKs p38, JNK, and ERK12 had been constitutively phosphorylated in RAW 267. 4 cells. how ever, only ERK12 phosphorylation was considerably accelerated in response to IL 32a stimulation. read the article This observation corroborated the truth that the addition of inhibitors for ERK12 and NF B suppressed every phos phorylation and consequently canceled IL 32a induced upre gulation of TNFa with the mRNA degree. Offered the delayed phosphorylation of I B and ERK12 beginning at thirty minutes within this research, IL 32a in RAW 267. four cells may not straight activate I B or ERK12. As an alternative, other molecules may perform a crucial purpose in I B or ERK12 activation of IL 32a, or the IL 32a TNFa axis may well use the MyD88 independent pathway reportedly related with the late inflammatory response of TLR 4.
Our examine also exposed that IL 32a induced IL 6 and MIP 2 at the same time as TNFa, but their induction was not canceled by inhibitors for NF B or MAPKs. This observation signifies that a signaling pathway other than NF kB or MAPKs could be concerned in IL 6 and MIP two expressions. IL 32g Tg mice obtained by using a promoter similar to that on the existing review reportedly exhibited no inhibitor OSI-930 apparent phenotype, but as soon as inflammatory colitis was induced with dextran sodium sulfate within the Tg mice, significant colitis occurred within 4 days. Curiosity ingly, at greater than 6 days just after DSS challenge, the degree of colonic irritation from the Tg mice was sig nificantly lowered, and recovery was additional quick than that in Wt mice due to the fact of elevated IL 10 levels in serum. In yet another research, IL 32b was reported to professional mote the manufacturing of IL ten in human cell lines. In accordance with our data on RAW 264. 7 cells, the level of TNFa in culture media peaked at twelve hrs soon after stimu lation with IL 32a and slowly decreased thereafter, whereas IL ten ranges greater from 24 to 96 hours right after stimulation.
In therapies containing both HP and TGF B1, the bio mechanical be
In treatment options containing the two HP and TGF B1, the bio mechanical advantages of HP have been dominated by TGF B1. Former do the job with articular chondrocytes stimulated by HP via the regimen utilised here demonstrated that the ERK12 pathway is needed for tensile home increase ment. Inhibition of ERK12 by U0126 blocked the tensile modulus enhancement observed with HP stimula tion. TGF B1 has also been proven to activate matrix professional duction in articular chondrocytes by way of ERK12. Inside the mixed HPTGF B1 treatment, the collagen and GAG contents and mechanical properties showed no major differences from TGF B1 remedy alone. In addition, no considerable distinctions have been observed amongst C ABC TGF B1 and full HPC ABCTGF B1 treatment in bio chemical content material or mechanical properties.
With both of these stimuli exhibiting action by the ERK12 pathway in articular chondrocytes, the result of TGF B1 could possibly be more robust on this cell population. Engineered costochondral cell neocartilage demon strated tensile properties that correlated with collagen content. In the present study, biomechanical, biophysical, and biochemical stimuli had been selleck employed with an objective of engineering robust tissues that might be capable of withstanding in vivo loads from cells that typically usually do not bear such loads. The results demonstrated that TGF B1 upregulated collagen synthesis related with increased tensile properties. In con trast, C ABC led to no alter in collagen synthesis about the cell level, however greater tensile properties by modula tion of fibril diameter and density.
The statistically signifi cant good correlation involving collagen written content per tissue wet fat and tensile stiffness and strength is so a perform of each collagen synthesis and fibril compaction. INK-128 Full HP C ABCTGF B1 treatment achieved 2. 2% collagenwet excess weight and also a tensile modulus of 2 MPa. A single may perhaps antici pate that even more efforts to boost collagen production, maturation, and organization will result in even more in creases in tensile properties of engineered tissues. Costochondral cells existing a clinically appropriate cell source that may be stimulated in vitro to create robust articular cartilage for use in load bearing joints. Costal cartilage may perhaps be isolated with ease surgically, and is un impacted by pathologies of the articulating joints, as well as arthritis.
Costochondral cells can be expanded in mono layer to increase cell quantity, and, in addition, chondro genic redifferentiation and self assembly result in a cell population that creates markers of articular cartilage kind II collagen, GAG, and SZP. While SZP gene and protein expression is absent in costal cartilage natively, engineered neocartilage demonstrated the pre sence of this protein, which functions in lubrication in load bearing, diarthrodial joints.
Fur thermore, inside the ER detrimental cell line MDA MB 231, E2
Fur thermore, from the ER damaging cell line MDA MB 231, E2 could not activate these kinases. The physiologic signifi cance of early ERK12 activation was confirmed by demonstrate ing that short pulse E2 remedy also stimulated cell proliferation. We and some others have confirmed that mER in breast cancer cells is responsible for this result by displaying that E2 peroxidase can stimulate proliferation and that this impact can be abolished by prior blocking of ER with ligand binding domain unique antibody. The inability to activate ERKs with E2 in an ER adverse cell line, plus the ability to undertake so in MCF 7 ER beneficial cells, was assigned by other people for the orphan G protein cou pled receptor GPR30. Nonetheless, current research with antisense GPR30 knockdowns unveiled that E2 stimulated MCF 7 cells proliferate also as cells with standard amounts of GPR30.
ERK activation connected with the capacity of those cells to respond to estrogens by proliferating as a result will not appear to get a function of GPR30 levels. Different levels of mER also established the E2 dose dependent phosphorylation of ERKs. Cells with lower amounts of mER responded to a restricted variety of concentrations. On the other hand, mERhigh cells mTOR inhibition responded to a very much wider range of E2 concentrations with a declining ERK activation at ten one hundred nmoll E2. This getting corresponds to our observation that ten nmoll and greater E2 decreases cell proliferation in mERhigh MCF seven cells, and it is constant together with the idea that E2 induces cAMP activated protein kinase A inhibition of MAPK pathways at these greater concentrations. Activation of ERKs continues to be linked to your proliferative cellular response.
Our accompanying kinase inhibitor Midostaurin report addresses other rapid estrogen elicited signaling responses that, in concert with ERK acti vation, can have an impact on and possibly balance cell proliferation responses. The classical antiestrogen ICI182,780 is effectively defined as an antagonist of the action of estrogen on the transcriptional level. In our scientific studies this compound was also a potent and quick ERK activator. With distinct kinetics, the transcriptionally inactive 17 estradiol also activated ERKs. It’s been demonstrated that pure antiestrogenER complex can bind to estrogen response aspects, but that the resultant transcriptional unit is inactive. This binding and inactivation is generally made use of for pharmacolog ical identification of ER participation in gene transcrip tion. However, within a yeast reporter technique antiestrogens induce ER dimerization and transcriptional action. As a result, whilst the published data disagree, it stays achievable that antiestrogenER complexes can exert speedy effects this kind of as induction of ERK phosphorylation along with other signaling results.
One particular way evaluation of variance with all the least sign
One particular way analysis of variance with all the least signifi cant difference post hoc test was applied to test for the differ ences inside the means of apoptosis rate. All P values are two tailed. P values of much less than 0. 05 had been thought of considerable. Outcomes Derlin 1 is overexpressed in the majority of human breast tumors The pathological diagnosis for all tumors was infiltrating breast carcinoma, having a tumor grade ranging from I to III. Immunohis tochemical analyses have been performed inside a blinded manner with respect for the pathology on the tissues becoming analyzed. The specificity of the primary antibody against human derlin 1 was validated. Whereas cytosolic staining was found to be strongly present in a breast cancer case, no staining was detected in sections in the similar sample when the section was topic to immunohistochemical evaluation working with the anti physique against derlin 1 that was pre incubated with peptide anti gen.
Derlin 1 signal intensity was graded as none to weak or as moderate to strong. From the 42 patients included within this study, 28 scored positively for expression of derlin 1, and ten of those 28 had really strong derlin 1 labeling. With respect to the quantity of cells labeled, greater than or equal to 25% labeling Mubritinib molecular weight was seen in all good circumstances. In addition, derlin 1 expression is predominantly present in the cytosol of tumor cells, but not in stromal cells. We also investigated derlin 1 expression in normal mammary glands. The staining intensity of standard mammary glands adja cent to tumor might be evaluated in 5 sections containing malignant tumors and typical glands in the similar slide.
Whereas no staining for derlin 1 was detected in the standard mammary glands, a signal of moderate or strong intensity was detected in all adjacent tumors. Furthermore, both IHC and Western blot analysis were employed to evaluate lev els of derlin 1 expression in yet another set of tumor samples with paired regular breast tissues from 13 sufferers. selleck inhibitor Among the 13 circumstances, only two typical breast tissues showed weak expression of derlin 1, whereas the other 11 typical breast tissues showed negative expression. Nonetheless, derlin 1 was charac terized by moderate or sturdy intensity in eight of 13 paired tumor samples. A representative Western blot evaluation is shown in Figure 3. Altogether, among the evaluated typical mammary glands, only two of 18 instances showed weak expression of derlin 1, whereas the other individuals showed adverse expression.
Derlin 1 expression correlates with tumor grade and lymph node metastasis For the girls diagnosed with breast cancer, we had information for tumor traits and noted no association among age, tumor size, and derlin 1 expression. Twenty seven of 42 females had grade 3 tumors. Twenty certainly one of the grade 3 tumors showed moderate or strong derlin 1 inten sity, whereas 7 of 15 grade 1 and grade 2 tumors have been derlin 1 positive.
Examination with the nucleotide modifications that led to STAT1 S
Examination of your nucleotide modifications that led to STAT1 STAT3 discriminating hpdODN B showed that they are compatible with prior in vitro DNA binding research, which include the preference for T at 1003 and 1005, dC at 1010 and dA at 1015 of STAT3. The fact that T at 1003 does not favor STAT1 binding is also in agreement together with the earlier suggestion that selection for any dG,dC base pair at position 7 is most likely to involve selleckchem Glu 421 which can accept hydrogen bonds from guanine within the minor groove. This has also been noted by others. Ultimately, altered recogni tion by a TF following single nucleotide modifications has been previously shown, for instance with NF B subunit recognition of B. One notable property in the hpdODN B is its dissymmetry. A symmetric version was tested and is appar ently not different from hpdODN B.
Intri guingly, although the preference of hpdODN D for STAT1 was anticipated from preceding data displaying its STAT1 precise binding, its basis isn’t clear and may well rest upon properties selleck chemical beyond nucleotide sequence including DNA shape. The shape and flexibility of DNA strands are identified to be influenced by their nucleotide content, here the 8 pyrimidine stretch in hpdODN B may perhaps confer a greater flexibility than hpdODN A and might account for any differential interaction with STAT3 Arg 423 and STAT1 Glu 421. In reality, the molecular dynamics research which describe a scissor like molecular movement upon DNA binding for STAT3, but not for STAT1 recommend that the flexibility on the DNA tar get may perhaps play a role in binding and consequently underly the preference of hpdODN B for STAT3.
It might also account for the higher sensitivity of STAT3 to an intact palindromic structure when compared with STAT1, as pre viously stated. Protein binding itself can impact DNA bending, as shown together with the high affinity target with the papillomavirus 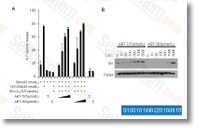 E2. Nevertheless, regardless of its effi ciency, the precise mechanism whereby the hpdODN B discriminates involving STAT1 and STAT3 in cells is not understood. Alterations in DNA shape could play a function within the preferential recognition of hpdODN B by STAT3, co aspects could also be involved in DNA recognition by STAT3, and may associate more efficiently when hpdODN B is applied. The procedure could possibly also be much more complex than mere differential DNA binding, STAT1 and STAT3 are reciprocally regulated as well as the relative abundance of their active forms may well itself play a important part in biological responses, as previously discussed. One more degree of complexity arises from the truth that in cells in which STAT3 has been suppressed, IFNg activated STAT1 induces the expression of mito genic STAT3 targets.
E2. Nevertheless, regardless of its effi ciency, the precise mechanism whereby the hpdODN B discriminates involving STAT1 and STAT3 in cells is not understood. Alterations in DNA shape could play a function within the preferential recognition of hpdODN B by STAT3, co aspects could also be involved in DNA recognition by STAT3, and may associate more efficiently when hpdODN B is applied. The procedure could possibly also be much more complex than mere differential DNA binding, STAT1 and STAT3 are reciprocally regulated as well as the relative abundance of their active forms may well itself play a important part in biological responses, as previously discussed. One more degree of complexity arises from the truth that in cells in which STAT3 has been suppressed, IFNg activated STAT1 induces the expression of mito genic STAT3 targets.
Not too long ago, GBMs have undergone a big scale muta tion scree
Not too long ago, GBMs have undergone a big scale muta tion screen and the molecular targets for this cancer is usually re evaluated. Essential to this strategy may be the identification of altered proteins or pathways that initi ate and or promote tumor development. Ideally, these mole cular targets are unique towards the tumor cell, and therapy precise to the alteration will not harm regular cells. You can find some very well-known genes mutated in GBM which include the tumor suppressors p53 and PTEN, and amplification or mutation with the EGFR and PDGFRA oncogenes. Unfortunately, molecular targeting efforts in GBM so far haven’t been translated into clin ical achievement, in spite of some promising results of targeted therapy in a few other cancers.
Even though there are several probable reasons why mole selleck cular targeting has not yet been productive in GBM, it can be doable that various or extra molecular targets in mixture will have far better good results. A recent survey of your coding sequence of 20,661 genes in GBM genomes has implicated lots of new mutated genes. Related to other cancers there are plenty of mutated genes in GBM and these genes cluster into essential pathways or gene groups. This clustering happens greater than possibility pre dicts, suggesting that these are a smaller variety of key cellular processes that must be altered inside the majority GBMs. One particular cluster of mutated genes reported by Par sons et al. was the ion channel genes. From the 555 genes involved in sodium, potassium, calcium and also other ion transport, 55 mutations had been detected affecting 90% in the samples studied with at least one particular somatic muta tion.
The statistical significance of this observation more info here was estimated to be p 0. 001 and the ion channels have been ranked as one of the leading gene clusters implicated by acquired mutations in GBM. Ion channels kind a important portion of cellular machinery and are accountable for transporting important ions across cell membranes, sustaining cell shape, cell volume and plasma membrane possible. Recent proof sug gests a role for ion channels in cancer progression and metastasis. Ion channels, for example sodium channels, potassium channels and calcium channels, have already been implicated for their role within a variety of unique can cers like colon cancer, prostate cancer, breast cancer and lung cancer. For example, the up regulation of voltage gated sodium channels is connected with pro gression of breast cancer metastasis. In this study, we report a correlation among ion channel mutations and patient survival. Twenty a single GBM individuals exactly where sodium, potassium and calcium channel gene sequences were known had been analyzed further for this study. GBM sufferers with a mutation in any with the sodium channel genes had a considerably shorter survival in comparison with those with wild kind sequence.
Although the detail protein protein interactions amongst TLR4, c
Though the detail protein protein interactions among TLR4, c Src, and p47phox are not identified, our benefits are the initial time to show a novel part of TLR4 MyD88 c Src p47phox complicated for mation in LPS induced NADPH oxidase activation and ROS production in HRMCs. Inside the future, we are going to fur ther ascertain which domains of TLR4, MyD88, c Src, and p47phox are involved in protein protein interac tions brought on by LPS. The MAPKs regulate diverse cellular applications by relay ing extracellular signals to intracellular responses. In mammals, you will discover more than a dozen MAPK enzymes that coordinately regulate cell proliferation, differentiation, motility, and survival. The top recognized will be the conven tional MAPKs, which include things like the extracellular signal regulated kinases 1 and two, c Jun amino terminal kinases 1 to three, p38, and ERK5 families.
MAPKs also happen to be shown to regulate VCAM 1 induction. Moreover, this is confirmed by our observation that LPS purchase MDV3100 induced VCAM 1 expression was lowered by inhibition of p38 MAPK, JNK1 two, or p42 p44 MAPK. ROS have been shown to stimulate p38 MAPK activation. Within this study, we demonstrated that LPS stimulated p38 MAPK, but not p42 p44 MAPK or JNK1 two activation was mediated by way of NADPH oxi dase ROS in HRMCs. Thus, we recommended that p38 MAPK mostly plays a crucial function in LPS induced NADPH oxidase ROS dependent VCAM 1 expression. AP 1 proteins are implicated in the regulation of numerous cellular processes such as proliferation and survival, differentiation, development, apoptosis, cell migration, and transformation.
AP 1 refers to a mixture of dimers formed in between mem bers of your Jun, Fos, and ATF families. their explanation Additionally, p38 MAPK has been shown to mediate ATF2 phosphorylation. Here, we showed that LPS markedly induced ATF2 activation, which was lowered by p38 MAPK inhibition. Hence, we demonstrated that LPS induced VCAM 1 ex pression through ROS p38 MAPK ATF2 in HRMCs. The transcriptional coactivator p300 is usually a ubiquitous nuclear phosphoprotein and transcriptional cofactor with intrinsic acetyltransferase activity. p300 controls the expression of various genes inside a cell type and signal particular manner, and plays a pivotal function in cellu lar proliferation, apoptosis, and embryogenesis. By catalyzing acetylation of histones and transcription fac tors, p300 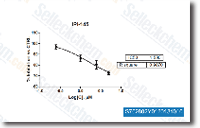 plays a substantial function in epigenetic regula tion. Current proof suggests that abnormal p300 function is related with deregulated target gene ex pression, and is implicated in inflammation. This really is confirmed by our observation that LPS induced VCAM 1 expression was decreased by inhibition of p300. Additionally, LPS straight stimulated p300 phosphoryl ation and the formation of ATF2 p300 complex via c Src ROS p38 MAPK.
plays a substantial function in epigenetic regula tion. Current proof suggests that abnormal p300 function is related with deregulated target gene ex pression, and is implicated in inflammation. This really is confirmed by our observation that LPS induced VCAM 1 expression was decreased by inhibition of p300. Additionally, LPS straight stimulated p300 phosphoryl ation and the formation of ATF2 p300 complex via c Src ROS p38 MAPK.
